All new findings of the State Key Laboratory of Wuhan University and Wuhan Virus Research Center
Recently, Zhao Haiyan's team from the State Key Laboratory of Virology of Wuhan University and Deng Zengqin's team from the Chinese Academy of Sciences Wuhan Institute of Virology jointly published a research paper entitled Structural and functional insights into the module of T cell collaboration by monkeypox virus protein M2 on Nature Communications. This study analyzed the complex structure of the first M2 protein of monkeypox virus with human B7.1 and B7.2, revealing that the M2 protein of monkeypox virus antagonizes the CD28/CTLA4-B7.1/2 signaling pathway by binding to B7.1/2 co stimulatory molecules, inhibiting a new immune escape mechanism activated by T cells, and providing new ideas for the defense of smallpox virus.
Monkeypox is a zoonotic disease caused by monkeypox virus infection. Since May 2022, this infectious disease has rapidly spread in multiple countries and has since been declared an international public health emergency by the World Health Organization. Although monkeypox no longer constitutes an international public health emergency since May 2023, its infection cases continue to increase. In July 2023, China reported 491 new confirmed cases of monkeypox. MPXV belongs to the family Poxviridae and the genus Orthopoxvirus. It is a brick shaped or oval shaped, enveloped DNA virus that also includes poxviruses such as cowpox virus and smallpox virus that can infect humans. The genome of Orthopoxvirus is approximately 200 kb, encoding and expressing up to 200 viral proteins, among which multiple proteins are involved in host cell interactions, regulating cellular metabolism and immune responses. Sequencing of the genes of the newly discovered monkeypox virus in May 2022 revealed large fragments of gene recombination in some strains, including an increase in the copy number of the M2 gene from the immune escape superfamily of the monkeypox virus, suggesting that the M2 protein may play a crucial role in the new transmission and pathogenic characteristics of the monkeypox virus.
In the classic T cell activation signaling pathway, the B7 ligands B7.1 and B7.2 on the surface of antigen-presenting cells bind to T cell surface receptors CD28 or CTLA4, playing a crucial role in regulating T cell activation and depletion. Multiple infectious pathogens and tumors can regulate the B7.1/2-CD28/CTLA4 signaling pathway to construct a suitable immunosuppressive microenvironment for better replication and survival.

In this study, the authors found that the early protein M2 secreted during the replication process of monkeypox virus and the M2 proteins of other smallpox viruses can specifically bind to B7.1 and B7.2 in the B7 family, without significant interaction with other proteins in the B7 family. In the high-resolution complex structure of MPXV M2-hB7.1/2, M2 exists in a cyclic form of homologous hexamers and heptamers, while hB7.1/2 binds to the outer side of the M2 protein ring in a 1:1 ratio.
Figure 1. MPXV M2 specific binding antigen presenting cell surface protein hB7.1/2
Structural analysis revealed a high overlap between the key binding regions of M2 on hB7.1/2 and the binding sites of CD28/CTLA4 on hB7.1/2, suggesting that M2 can competitively inhibit the interaction between CD28/CTLA4 and hB7.1/2. Cell and competitive BLI experiments further demonstrate that M2 binding to hB7.1/2 can block the interaction between CD28/CTLA4 and hB7.1/2. Sequence alignment revealed that the interaction imprint on M2 is highly conserved in the MPXV branch and other orthopoxviruses, indicating that M2 may mediate a conserved immune escape mechanism by blocking the interaction between B7 ligands and CD28 family receptors by binding to hB7.1/2, thereby inhibiting T cell activation. Interestingly, in vitro T cell activation experiments have found that the polymeric form of M2 can effectively inhibit the co stimulatory pathway mediated T cell activation of hB7.1/2, while the affinity between monomer M2 and hB7.1/2 significantly weakens, and monomer M2 almost loses its ability to inhibit T cell activation, indicating that the high affinity binding property of aggregate form M2 with hB7.1/2 plays a key role in virus evasion of T cell immune response.
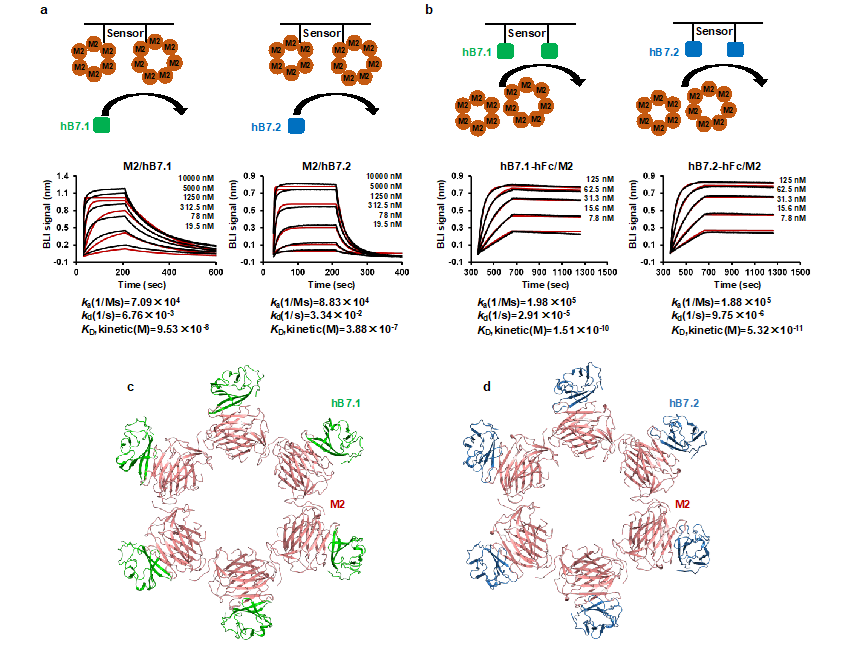
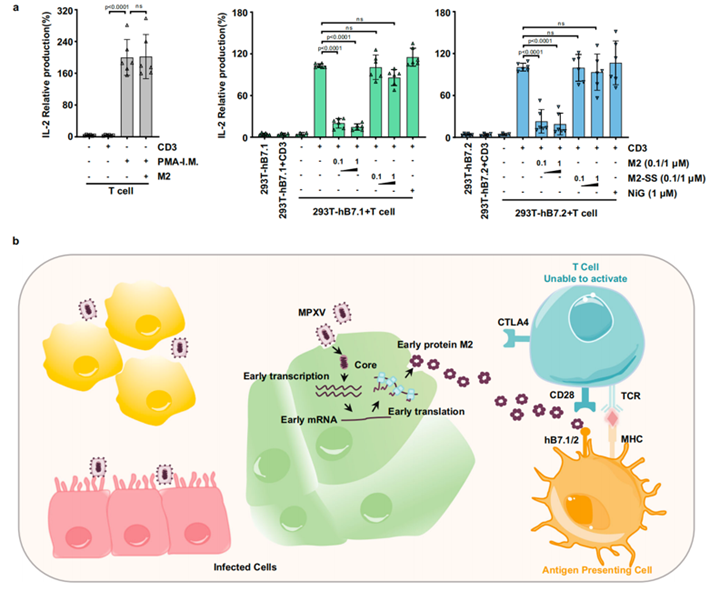
Figure 2. MPXV M2 inhibits T cell activation
In summary, this study for the first time analyzed the structure of the M2 protein of Orthopoxvirus, revealing that the highly conserved immune escape protein M2 in Orthopoxvirus enhances its binding affinity with the T cell co activator hB7.1/2 by forming a polymer, effectively blocking the interaction between CD28 and hB7.1/2, thereby inhibiting the viral immune escape mechanism of T cell activation.
Zhao Haiyan, a researcher from the State Key Laboratory of Virology of Wuhan University, and Deng Zengqin, a researcher from the Chinese Academy of Sciences Wuhan Institute of Virology, are the co corresponding authors of this paper. Yang Shangyu, a master's student from the School of Life Sciences of Wuhan University, Yu Feiyang, a doctoral student, and Wang Yong, a doctoral student from the Wuhan Institute of Virology, Chinese Academy of Sciences, were the co first authors of the paper. The research was supported by the National Key Research and Development Program, the National Natural Science Foundation of China and the Chinese Academy of Sciences' First Action Talent Introduction Program.